Food Microbiology
oCelloScope in dairy mold research
Yoghurt is a difficult matrix to analyze. Casein in yoghurt reflects all light and makes it an impossible assay in microscopy. Today, most food companies study the growth and inhibition of molds using classical agar plates and rely on visual inspection. Analysis time ranges from 3 to 21 days depending on temperature.
BioSense Solutions have developed an assay to study growth of molds in yoghurt. We give the user the ability to both count spores in samples and quantify germination and overall growth. Analysis time is just 1 to 5 days depending on temperature.
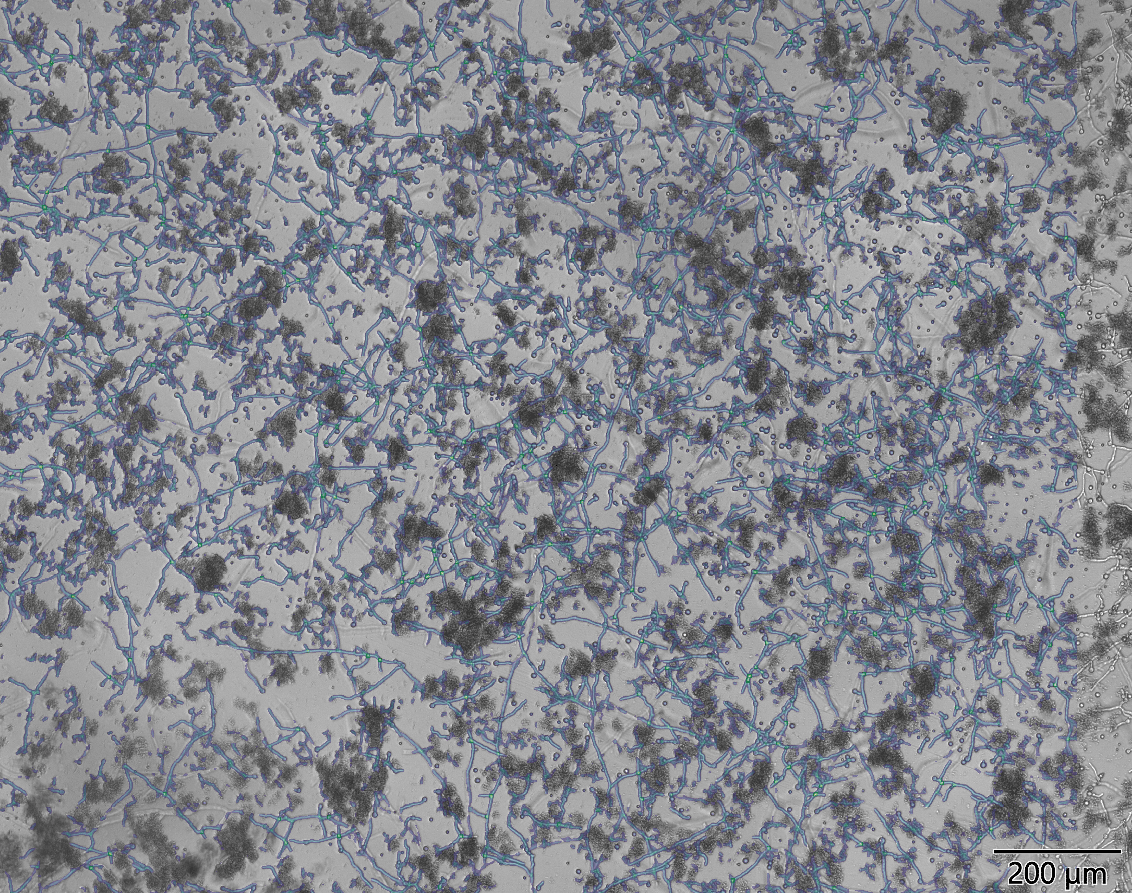
On the left is a sample of yoghurt inoculated with Penicillium spores. Dark elements is casein. Mycelia is tracked with blue overlay and in green branching/crossing is marked. We are able to compare and quantify growth of different molds or treatments.
The assay is most likely beneficial in any food dairy application where the matrix can be homogenized.
oCelloScope Yeast
Investigating Live-Dead status in a yeast population is often a manual and uncertain process in both R&D and production. On the right is a video of how we work with yeast.
We have stained a yeast sample to investigate, if what appears dead is dead, and if what appears alive will actually grow. With the oCelloScope we can measure growth, count, proportionate, and validate yeast cultures.
We use time-lapse imaging and algorithms to help automate and simplify processes.
oCelloScope in probiotics discovery
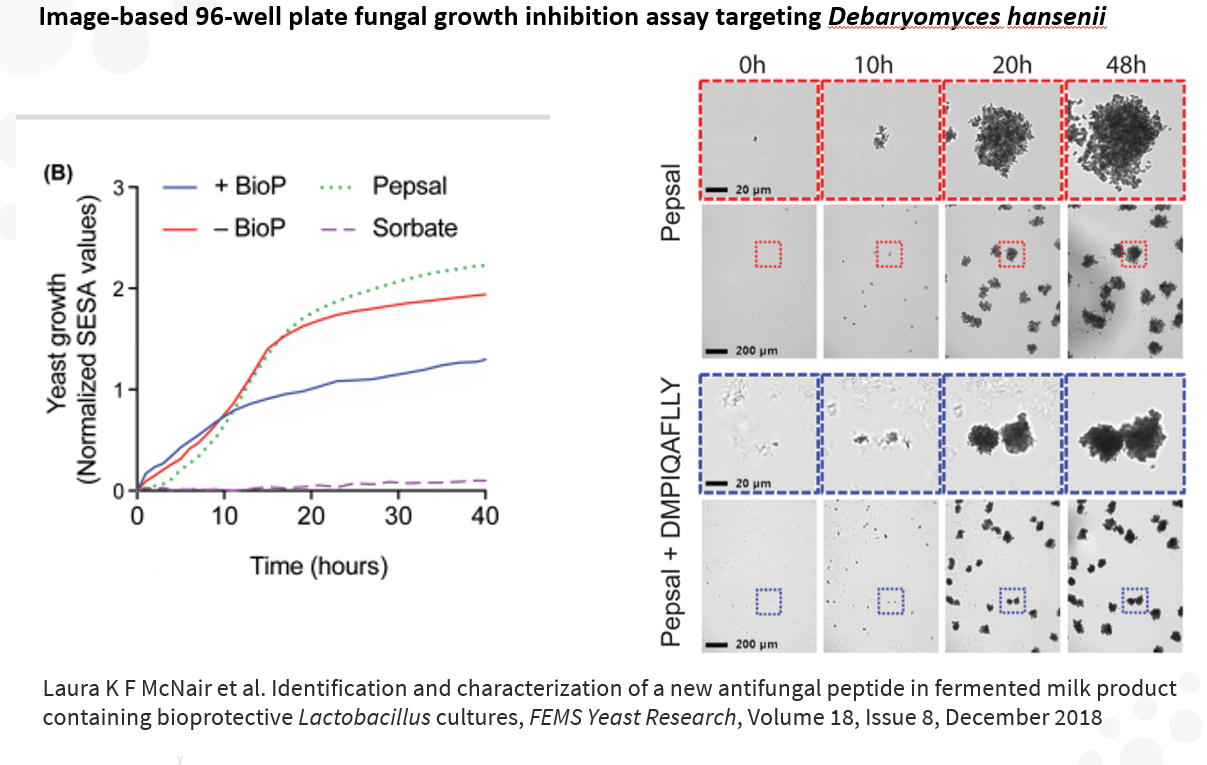
Left: Aqueous samples of sour cream produced by fermentation of milk and starter culture with the addition of bioprotective strains (+BioP) compared to those without (–BioP).The growth was compared to growth in pure saline peptone (Pepsal) and a control in saline peptone
Right: Debaryomyces hansenii grown in pepsal containing peptide exhibits altered morphology. Representative images from the oCelloScope 96-well plate fungal growth inhibition assay.
oCelloScope in food mold discovery
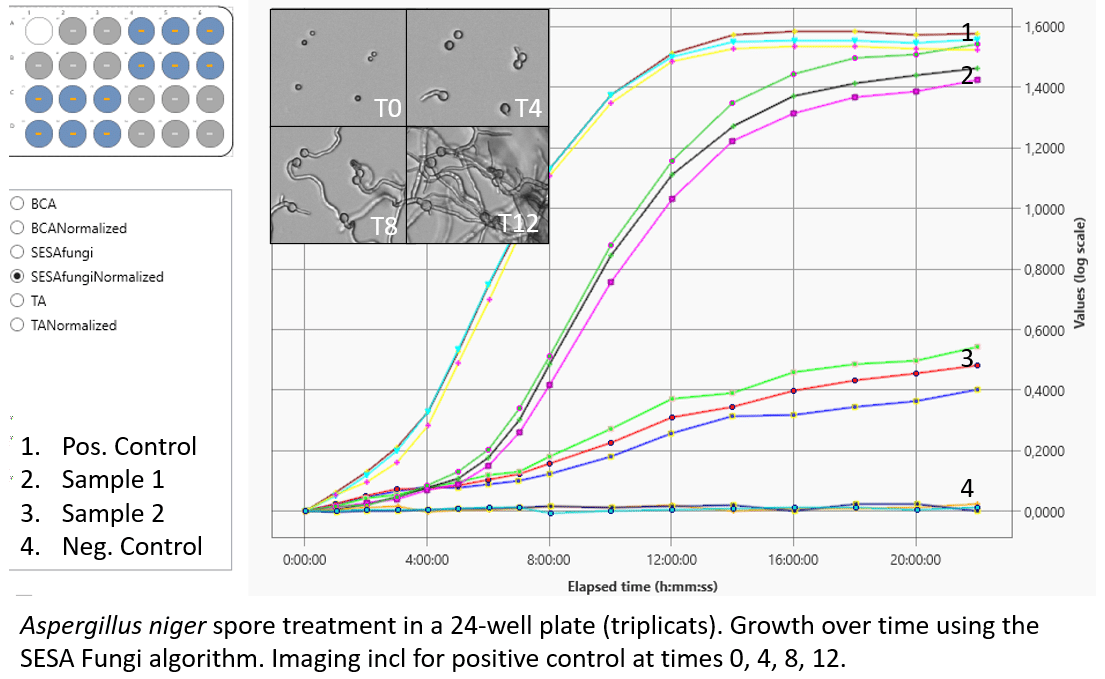
Aspergillus niger, or black mold, is a common fungus that appears on decomposing starchy fruits and vegetables as well as on damp walls as a component of mildew. Many of our users study Aspergillus niger and germination of spores. In the view on the right you see a snip from the UniExplorer software showing the growth kinetics of germinating spores.
Note, that we are using the SESA Fungi algorithm which detects not only area of an organism with a dark outline but also measure area of brighter/transparent organisms such as fungi. The algorithm makes the platform superior to using a plate-reader.
Preservatives testing in 384 well multi-well plates.
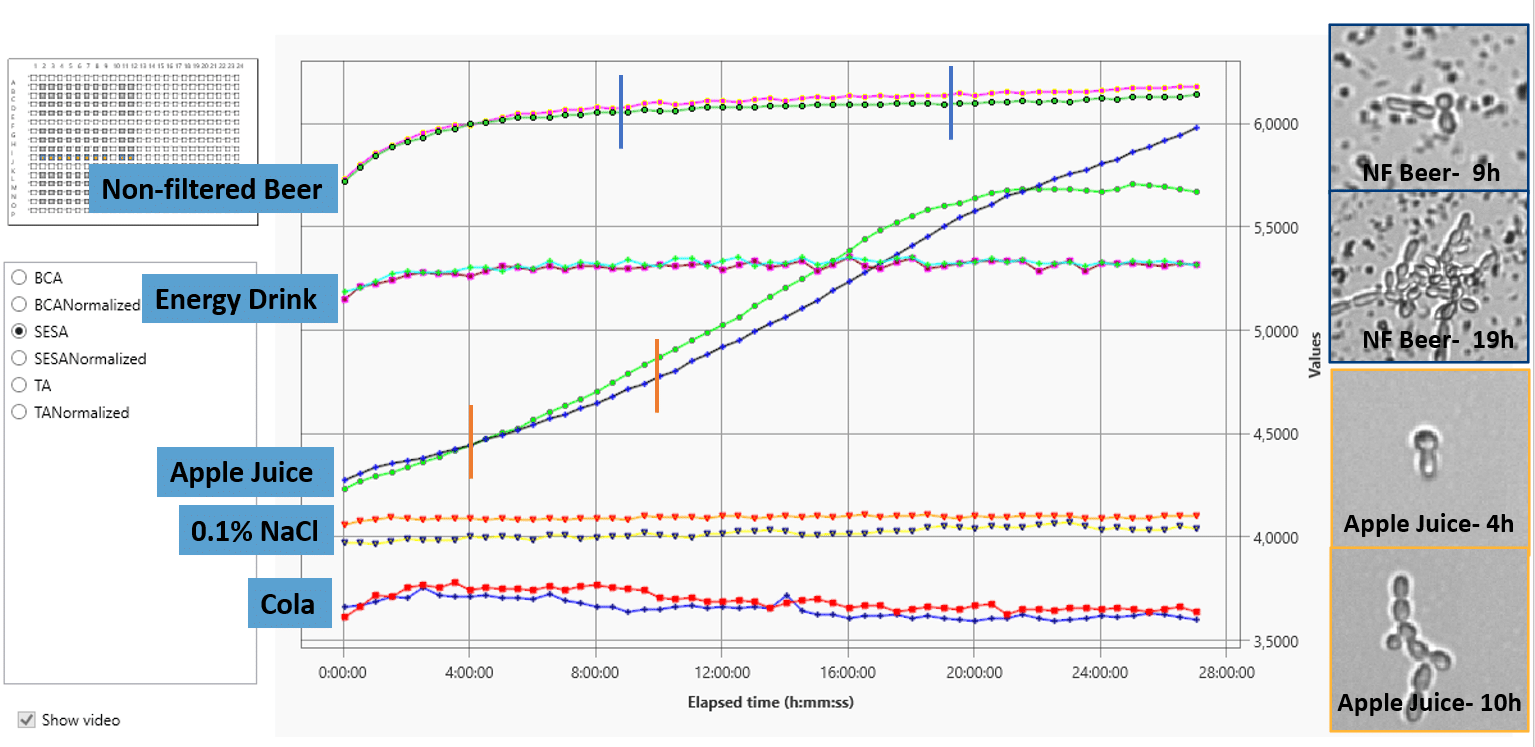
Left: yeast spike experiment in 5 different matrices (Cola, Apple Juice, Non-filtered Beer, Energy drink and 0.1% NaCl). Samples were inoculated with yeast and experimpent set to run for 27 hours with imaging of wells every 30 minutes. Growth kinetics module was chosen and shows growth of yeast in Apple Juice and Non-filtered beer. No growth was detected in Cola, Energy drink or 0.1% NaCl. Time point images of growth are inserted for non-filtered beer and apple juice. Note, the fast detection time in the automated system.
The oCelloScope can be used to test new preservatives against fungi, yeast and bacteria in high throughput. In addition, testing quarantined batches of beverages can lead to a faster release of products and thus a longer shelf life.
Analyse complex samples and discriminate between different morphology
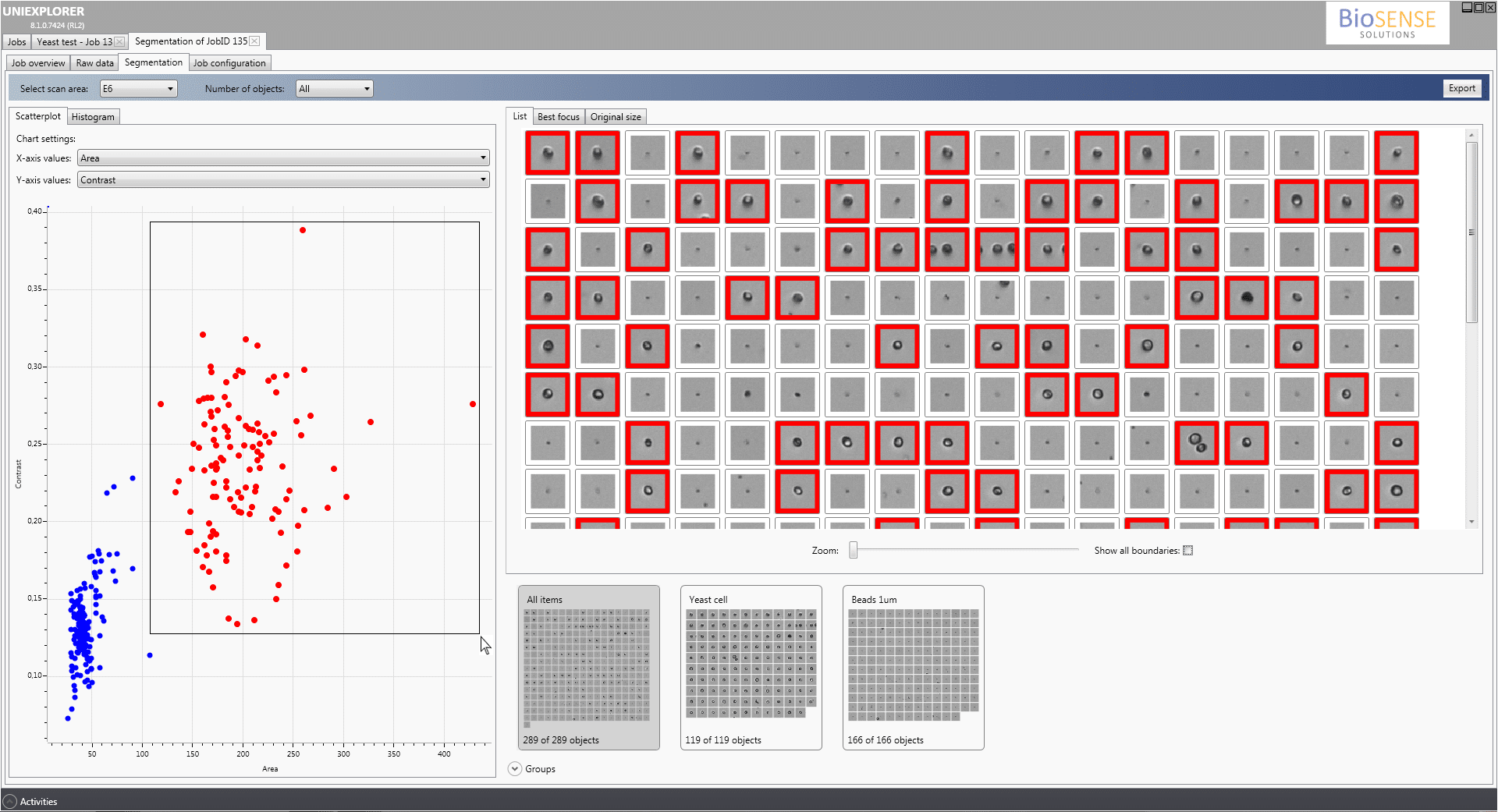
With the build-in segmentation algorithms, the UniExplorer software enables the user to automatically perform single cell analysis and discriminate and characterize the dynamics of different cell types in a complex sample.
More than 20 morphological features, including cell size and shape factors of each single object can be calculated and visualized in scatter plots and histograms, enabling the user to group the desired objects and analyse the morphology changes over time of the desired group.
As an example, discrimination between vegetative cells and spores in fungi or bacterial sporulation experiments can be performed.
Ready to get started?
We are ready to help you solve your tough research challenges. Our highly qualified team can help advance your next big discovery.