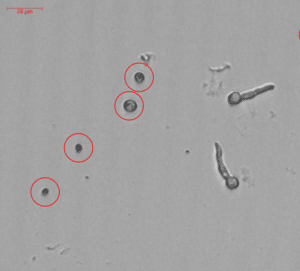
Counting spores in fungal samples
Fungal cell counts are used in both R&D and production settings in a wide variety of applications. Most cell counts are based on manual microscopy and this we aim to change. BioSense Solutions employ machine learning algorithms to count spores. Spores can even be counted in dirty samples containing hyphae, fibres or other debris. If work is performed in micro titer plates, and we know the sample volume, spores/ml can be automatically calculated. Counts can be performed as end-point or used in time-lapse studies to quantify germination.
The oCelloScope is routinely used to count final products in both the brewing industry and in global agriculture (Bio pest control).
Even distribution of cells is paramount in getting correct results. It is well-established that cells can populate the edges of a well and thereby affect count. Literature finds that edge effects are minimized if sample and plate have same temperature. In addition, pipetting into wells can affect distribution.
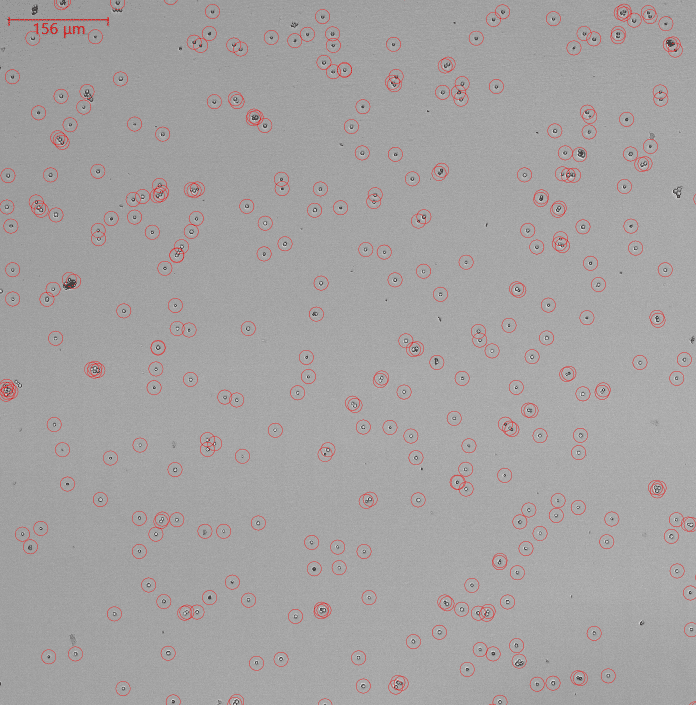

End-point SporeCount
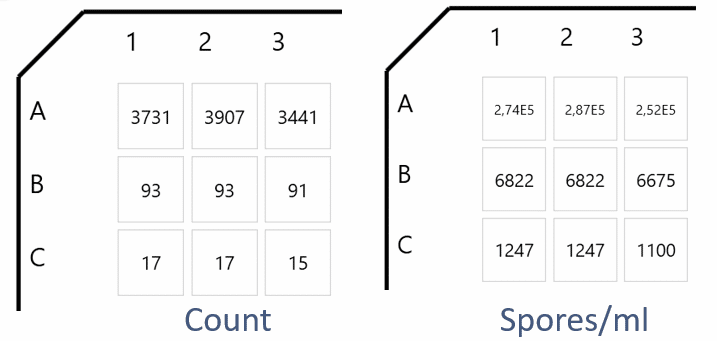
With an even distribution of cells, triplicates will yield similar results. In the software, wells are shown and findings can be displayed as actual number or spores/ml. Spores/ml is calculated from sample volume.
Note that when you inoculate 100µl sample into a well of a 96-well plate, spores will settle at different rates. To get precision in your count it is advised to wait 1h before the SporeCount analysis is run. It is advised to increase number of images in wells from 10 to 70 to capture a larger area in wells. In addition, it is given that plates are quality manufactured and have an even surface.
Triplicate counts of spores in a 96-well plate is visualized on the left – both as actual number and spores/ml.
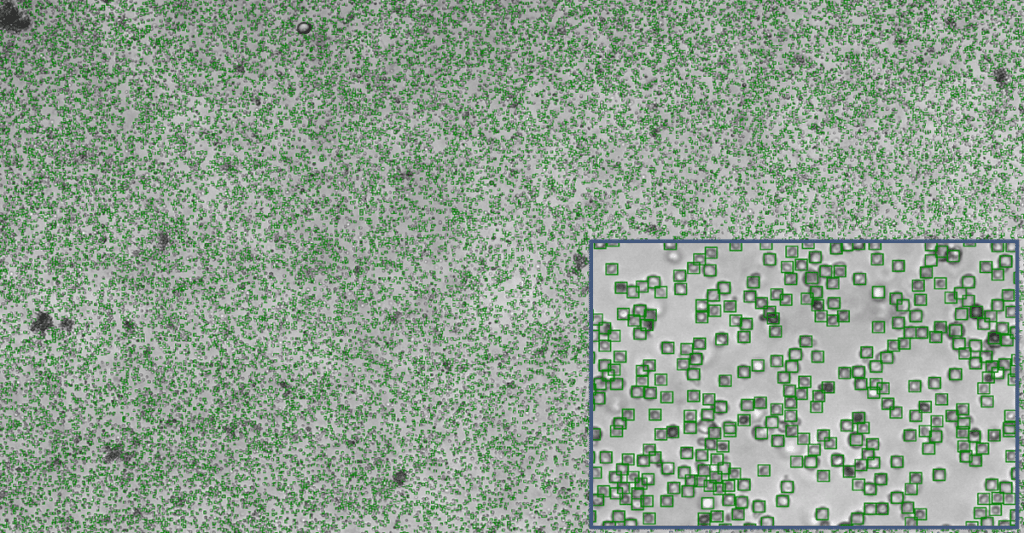
Yeast SporeCount
The deep learning algorithm for yeast is designed to count yeast cells even up to high concentrations. In image on the right you see an near to perfect count of yeast. In any case, this is much better than manual counts and what we have been seen from competing technologies. Counts can be run as endpoint or time-lapse in any flat bottomed microtiter plate format.

Time-Lapse SporeCount
A common application using the SporeCount module is to time-lapse your spore development. Spores will be tracked until the time-point where they develop a germ-tube. What you get is exact time of germination and you are able to easily quantify germination. Resulting curve on the left can be set to show actual spore count or percentage. When set to percentage you can quantify how many spores in a population will germinate.
Note how curve increase from starting point. This is due to settling of spores in wells. This can take hours depending on fungal species.
From Dormant to Isotropic growth
The SporeCount module is based on Deep Learning algorithms. Algorithms are designed to track spores from dormant to the point where they develop a germ-tube. Algorithms are species specific for highest level of precision. As fungal spores comes in a variety of shapes, we offer to custom build deep learning algorithms for your species – contact us to learn more.